Circulating tumor DNA (ctDNA) is a kind of free DNA released by tumor cells after necrosis and apoptosis. Because it circulates in the blood, the detection of ctDNA can reflect the tumor state in time. In 1940, Mandel and Métais first discovered the presence of free cellular nucleic acids in the blood of healthy people. Later, Leon et al. found that the circulating free DNA (cfDNA) concentration in the blood circulation of tumor patients was higher in healthy people. The cfDNA is derived from tumor, which we call ctDNA.
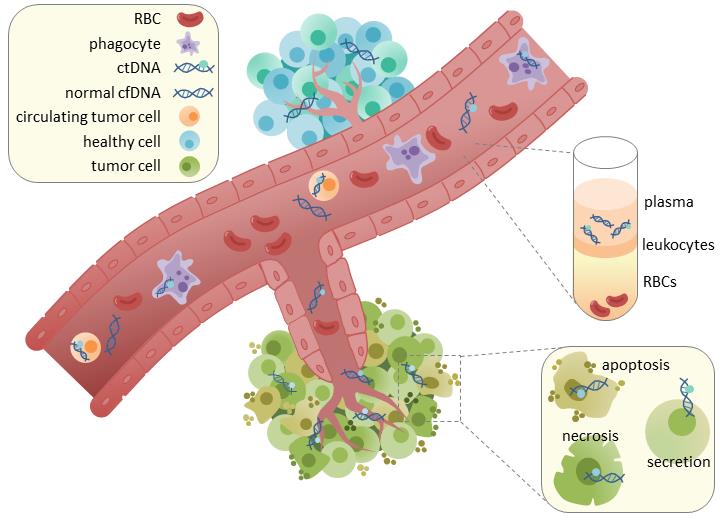
Source of CtDNA
Through recent studies, it is speculated that the large amount of DNA present in circulating blood of tumor patients may be derived from the following pathways: 1 Apoptosis of tumor cells. As the tumor volume increases, the apoptotic cells also increase in the normal physiological environment, and the scavenger cells will clear the apoptotic cells in time. However, scavenger cells cannot effectively eliminate the rapidly increasing apoptotic cells, which inevitably lead to the release of cellular contents into the blood circulation, including ctDNA. In fact, most ctDNA fragments are 180 to 200 bp in length, suggesting that most of the ctDNA in the circulation is the amount of passive release of ctDNA from the apoptotic cells to the blood circulation, depending on the location, size, and vascular distribution of the tumor. 2. The hypothesis of cell necrosis and autophagy. CtDNA may also be derived from necrosis, autophagy, necrotic apoptosis, and other micro-environmental pressure and chemotherapeutic-induced physiological activities. Unlike apoptosis, necrosis is a process of disease that randomly and incompletely digests genomic DNA and produces larger DNA fragments. 3. Tumor cells are lysed in circulating blood or micrometastases. The ctDNA may be a lysate of circulating tumor cells or tumor micrometastases. 4. Tumor cells continuously release DNA into the blood circulation. The hypothesis is that cells can actively release DNA into the bloodstream in the form of DNA-nucleoprotein complexes.
Application of ctDNA in tumor therapy
A common method for clinically detecting tumor status is tissue biopsy. However, due to the heterogeneity of the tumor, a single region of tumor biopsy does not fully understand the mutation profile of the entire tumor. In addition, since biopsy cannot be performed continuously, it is impossible to understand the progress of the tumor and the changes in the genome during the treatment in real time. Mutations by ctDNA can broadly represent changes in the tumor genome and provide a non-invasive method of tumor sampling that overcomes many of the limitations of current biopsy. In addition, ctDNA testing has shown great potential for early tumor diagnosis and provides a new direction for controlling cancer in the early treatable stage. Although ctDNA detection has great potential for development, its application to the clinic still faces many obstacles that need to be overcome. First, there are no specific operational standards for each step of purifying cfDNA from specimens to detect tumor hotspot gene mutations. In addition, although the methods for detecting ctDNA are constantly evolving, the cost and practicality of clinically applicable will be subjected to a severe test. Currently the most sensitive technology, such as, dPCR, etc. can only be used for hot spot mutation detection and cannot be applied to mutation discovery. Only by first sequencing the biopsy genome, and designing highly specific probes based on the information of the high frequency mutation sites, the dPCR method can be used for ctDNA detection. Although the NGS method can perform full exome sequencing on patients, the large genome makes sequencing cost very high, which limits its clinical application. Therefore, these methods can be applied clinically as early as possible only if the ctDNA detection technology forms a uniform operating standard and further reduces the cost of testing.
