Introduction of circadian rhythm molecules
There are various rhythms in the life activities of nature. The most common of all rhythms is the circadian rhythm. Sleep not only has a circadian rhythm, but many other behaviors and physiological activities also have circadian rhythms. The study found that the circadian rhythm is inherently endogenous, controlled by the biological factors, and can adapt to changes in the environment (such as light, temperature, and eating time.); environmental changes can cause temporary confusion of circadian rhythm, thus affect the physiological activities of living organisms. From single-celled blue-green algae to multi-celled people, most of the life activities of various organisms have circadian rhythms and follow similar molecular regulatory mechanisms. Since the discovery of the first gene controlling circadian rhythms in Drosophila in 1971, the intrinsic regulation mechanism of circadian rhythms has become clear. Michael Rosbash and Jeffrey C. Hall of the University of Brunswick and Michael W. Young of Rockefeller University won the 2017 Nobel Prize in Physiology or Medicine for their achievements in the study of the molecular mechanisms of circadian rhythms.
Molecular Mechanism of Circadian Rhythm Regulation
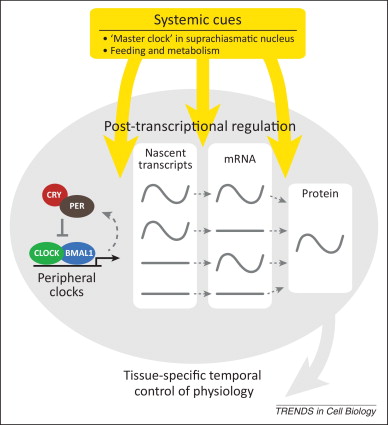
The regulatory mechanism of circadian rhythm is evolutionarily conservative, and mammals follow a similar regulatory mechanism as Drosophila. The early work of finding circadian rhythm regulating genes was mainly done in fruit flies. The first discovered circadian rhythm clock gene is per gene of Drosophila. Ronald Konopka and Seymour Benzer used the characteristics of the emergence and diurnal activity of Drosophila as indicators, and screened three mutant Drosophila strains with abnormal circadian rhythms: one had no rhythm, one rhythm cycle shortened to 19 hrs, one rhythm changed to 28 hrs. The three mutations are caused by different mutations of the same gene. They named the gene as period. Unrhythmically is per0, the short rhythm is pers (per short), and the rhythm is long per (per long). Further research found that the PER protein encoded by the per gene is a transcriptional regulator. In the further research, other biological clock genes regulating the circadian rhythm of fruit fly have been found, jrk, tim, cry, cyc, dbl and so on. The jrk gene product regulates the expression of the per gene and is also regulated by the PER protein. The protein encoded by the tim gene is regulated by light, and the product of the cry gene is affected by the binding to the PER protein. The CRY protein enters and exits the nucleus and mediates the effects of light on the inner circadian clock of the body by regulating the expression of the per gene and the tim gene. These circadian clock genes that regulate circadian rhythms found in Drosophila mostly have counterparts in the mammalian genome. The first gene regulating the mammalian circadian clock, tau, was discovered in 1988 and successfully cloned in 2000. Its gene product is a protein kinase CK1ε. The second discovered mammalian circadian clock gene is the clock protein gene, whose gene product CLK protein is a transcriptional regulator and is a homologous gene to the jrk gene of Drosophila. The first discovered per gene in Drosophila also found homologous genes in mice and humans in 1997. There are three per genes in mammals, per1, per2 and per3, and their expression products have circadian rhythm changes and are subject to regulation of the clock gene. Other circadian clock genes found in mammals include the cryptochrome genes cry1 and cry2, the brain and muscle aryl hydrocarbon receptor nuclear translocation protein gene bmal1, and the orphan nuclear receptor gene Rev. The basic pattern of circadian rhythm regulation of circadian clock genes is similar in various organisms: transcription-translation-negative feedback inhibits transcriptional patterns, circadian rhythmically expresses circadian clock protein as a regulatory factor, promotes expression of other related genes, and then inhibits its own transcription through negative feedback. The current study found that there are mainly two feedback loops in the regulation mechanism of mammalian circadian rhythm. In the first loop, the proteins mCLK and BMAL1 expressed by clock and bmal1 gene first form a heterodimer in the cytoplasm and then transport it into the nucleus. In nucleus, the heterodimer binds to the promoters of the cry and per genes, activates their transcription, and promotes the transcription of the circadian clock-related genes. The CRY and PER proteins are expressed after the initiation of the cry and per genes. As transcriptional translation proceeds, the CRY and PER proteins accumulated in the cytoplasm gradually increase. They are activated by phosphorylation and form heterodimers by interaction and transported to the nucleus. The heterodimers binds to the CLK/BMAL1 dimer by virtue of the PAS domain of the PER protein, which reduces the transcriptional activation of the CLK/BMAL1 dimer and achieves the inhibition of self-transcription. The PER protein, which has been phosphorylated in both the cytoplasm and the nucleus, is degraded after being re-phosphorylated by CK1ε kinase. In the second loop, the dimer CLK/BMAL1 activates the transcription of the orphan nuclear receptor gene rev, and its translation product, REV-ERBα, inhibits the transcription of the bmal1 gene by binding to the promoter of the bmal1 gene. Therefore, when mPERl and mPER2 are transported into the nucleus to form a stable negative regulatory polymer with mCRY1 and mCRY2, the transcriptional activity promoted by CLK/BMAL1 dimer can be directly inhibited; while inhibiting the expression of mPER1 and mPER2, it also inhibits the transcription of the rev gene, which causes the transcriptional inhibition of the bmal1 gene to be released, and then the expression of BMAL1 protein increases, and enters the next rhythmic oscillation cycle.
References: