Antibodies are one of the important reagents in life sciences and clinical medicine. Although antibodies have been widely used, there is a lack of established criteria to guide how they should be validated before use. As a result, many commercial antibodies have not been fully validated prior to use and/or lack of further confirmation resulting in failed or unreplicable results, even projects being abandoned, resulting in significant time, money and sample loss. Therefore, the establishment of antibody verification standards has become an urgent need.
Antibody validation consists primarily of the following components: demonstrating specificity (the ability of antibodies to distinguish between different antigens), proving affinity (intensity of antibody binding epitopes), and finally demonstrating reproducibility. However, although this definition of antibody validation is reasonable, there are still some problems with the widespread application or implementation of validation standards.
The task of verifying the antibody should have been done by the antibody supplier. However, although the seller is responsible for the quality of the reagents sold, antibody performance may be affected by a number of other factors. For example, antibodies may change during transport due to improper storage at low temperatures, or although they can be successfully validated in a general biologically relevant system, end users need to use it in their own experimental systems. Therefore, the end user also needs to perform secondary verification of the antibody.
By taking a few small steps, the researcher can effectively reduce the risk of the experiment in subsequent experiments. First, the researcher should be aware of the usage information provided in the product specification, including the application that has been validated for it, the appropriate protocol and the recommended dilution.
There are also different methods for antibody validation for specific applications, including Western blot (WB), immunohistochemistry (IHC), immunocytochemistry (ICC), immunofluorescence (IF), ELISA, immunoprecipitation (IP), chromatin immunoprecipitation. (ChIP) and peptide arrays, etc. In terms of verification, each assay has its advantages and disadvantages (Table 1).
Table 1. Advantages and disadvantages of several antibody validation methods.
| Assay | Advantage | Disadvantage |
| Western Blot | Easy and simple assay Ideal for denatured proteins | Difficult to optimize Time-consuming A small number of antibodies can be tested per run |
| ELISA | Quantitative assay High throughput | Does not determine if the antibody is specific or cross-react |
| IHC/ICC | Routinely available and relatively inexpensive | Difficult standardization Epitope accessibility can vary in fixed tissues Difficult to quantitate |
| IF | High throughput Easy to optimize | Does not determine if the antibody is specific or cross-reacts |
| siRNA knockdown | KD cell lines can be used in all assays (WB, ICC/IHC, Flow cytometry) | KD is transient Difficult to optimize-requires several siRNA sequences |
| Knockout cell lines and mouse models | Best negative controls, since they guarantee no expression of the target gene May be used in all assays (WB, ICC/IHC, Flow cytometry) | Cell lines for specific genes are not always available or lethal KO mouse models take over a year to develop and are often non-viable |
| MS | Confirms specificity High throughput | Requires use of mass spectrometer and trained personnel |
| SPR | Real-time analysis Label-free Highly sensitive | Requires immobilization Requires meticulous experimental design High sample consumption |
| MST | Rapid assay (KD in 10 min) Low sample consumption (pM/nM) and small volume (<4 ul) Immobilization free-in solution measurements Label-free (optional) | Requires specialized equipment |
Which method should be used and how several methods are used to validate antibodies depends not only on sensitivity, specificity and reproducibility, but also on the environment in which the particular antibody is to be used. This means that if a given antibody is only intended for WB, they only need to be tested in this case. However, if you need to use it in other assays, you should also test it again in the application environment. This helps save time and cost associated with the antibody validation process.
Specificity is a measure of the ability of an antibody to distinguish between different antigens. Even if the antibody may be sensitive to the target protein, it will show a lack of specificity due to cross-reactivity with other proteins. In this sense, it is a basic requirement for a good antibody to confirm that the antibody specifically binds to the target protein.
Several questions need to be identified before analyzing the specificity of a given antibody. The first is what are the types of immunogens that are used to produce this antibody? How is the target protein in the sample to be analyzed constructed? The first question is not easy to answer, especially if the antibody manufacturer does not disclose the type of immunogen. The reason for understanding this information is that the specificity of an antibody depends in part on whether the type of immunogen is a synthetic peptide or a purified protein. Since synthetic peptides do not necessarily summarize the 3-D structure or post-translational modification of native proteins, antibodies produced using synthetic peptides may not recognize native proteins. If the immunogen is a purified protein, the denatured protein is difficult to bind to the antibody. Thus, when the immunogen is a synthetic peptide, the antibody can be applied to WB but not to IP or IHC.
WB is considered to be the first step in the evaluation of new antibodies because it is the ideal assay for specific validation of denatured protein antibodies. However, in experimental assays where the antigen is in its natural conformation (eg, IHC), it is not the ideal standard method for antibody binding. Because antibodies may recognize a particular epitope in fresh tissue, but different epitopes are recognized in the fixed tissue, the method used to fixed tissue complicates the problem. This happens because unexposed epitopes in natural proteins become accessible after fixed processing and vice versa. If you only need to verify the recognition of the denatured antigen by the antibody or whether it is suitable for WB, then the assay can be used as a first step of verification. The result when the antibody is specific for the selected target is the observation of a band on a target of known molecular weight. If multiple bands are present, they may represent different post-translational modification states, decomposition products or splice variants, or may also be non-specific binding. Therefore, the results of these experiments may appear to be incapable of determining antibody specificity.
Another disadvantage of WB is its low detection throughput because it usually only tests one antibody at a time. However, recently, by converting WB to a capture format, a high-throughput detection method called PAGE-MAP was developed for antibody verification. In this method, a biotinylated protein sample is subjected to PAGE, and then the size-separated protein is incubated with a microsphere-based barcode antibody array for multiplex IP, and the captured protein is labeled with streptavidin for flow through Magnetic cytometry (ie, microsphere affinity) was performed for magnetic bead detection. In addition, size-separated proteins can also be used for parallel readout shotgun mass spectrometry (MS), which can be used to roughly estimate specificity and for selection of antibody target levels sufficient for immunoprecipitation. This new method is powerful, screens thousands of antibodies, recognizes antibodies that bind to the same protein, and provides a means for large-scale antibody validation.
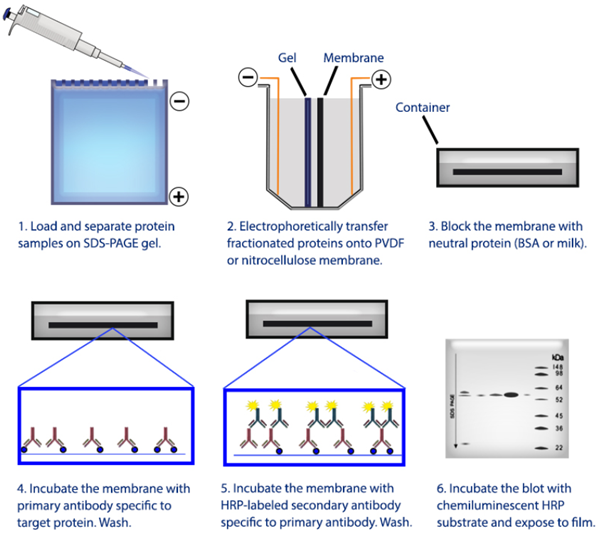 Figure 1. Overview of the western blotting procedure.
Figure 1. Overview of the western blotting procedure.
In addition to the WB method, the use of blocking peptides to assess antibody specificity is also a common approach, particularly for IHC. These peptides are identical to the peptides used to produce antibodies, and when over-incubated with them, they can be used as immunoneutralizing antibodies. In the validation experiment, the unneutralized antibody was used as a control to stain the sample tissue. If the antibody is specific, incubation with a blocking peptide will result in the disappearance of staining on the tissue. However, this assay has the disadvantage that it does not verify the selectivity of the antibody for the antigen, since the non-specific binding activity of the antibody will also be inhibited by the blocking peptide. Thus, blocking peptides can prove that antibodies are bad, but they do not prove that antibodies must be good.
In the process of antibody validation, the key to demonstrating antibody specificity is the correct use of controls. As in the antibody verification experiments on the cannabinoid CB2 receptor, it has been found that the conventional practice of using only a positive control to verify an antibody is insufficient to ensure the reliability of the antibody. In this validation study, although many techniques (eg, WB, mass spectrometry and blocking peptides) and excellent positive controls were used to obtain a number of positive results indicating the effectiveness of anti-CB2 antibodies. However, when knocking out a negative control to test antibodies, this antibody was found to be non-specific for the CB2 receptor protein. A similar situation may occur due to the similarity of relative epitope regions between proteins, or because antibodies have multiple epitopes. The best negative control is knockout cells/animals that do not express the protein of interest, and closely related proteins can also serve as good negative controls. The best positive control is a cell which non-expression the target protein and transfected with the protein of interest. If knocking out the cells is too difficult to obtain, then siRNA or shRNA knockout cells can be selected as controls. Although negative control is necessary, there is also a need for positive control.
Antibody binding affinity is another parameter that can be used for antibody validation. It refers to the strength of the binding of an antibody molecule to an epitope. It is usually reported as the equilibrium dissociation constant (KD), which is the ratio of the antibody dissociation rate or koff (the rate of dissociation from the antigen) to the antibody association rate or antibody binding rate Kon(the rate of binding to the antibody).
Affinity determination of monoclonal antibodies can be performed with high precision because they are selective only for the same epitope, but in the case of polyclonal antibodies, since the epitopes they detect are heterogeneous and are various A mixture of antibodies with different affinities. Therefore, only the average affinity can be obtained. In order to determine antibody affinity, the following methods have existed. These include ELISA-based methods, as well as other biophysical methods such as microscale thermophoresis (MST) and surface plasmon resonance (SPR).
ELISA is the most popular method for studying antibody affinity. They do not require the use of large amounts of antibodies and antigens, nor do they require purification of the protein. In this method, a fixed concentration of antibody is incubated with the antigen in solution until a steady state is reached. Then, the concentration of the unbound antibody was measured by an indirect ELISA. This method requires prior preliminary experiments to determine the concentration of antibody used within the linear range of the ELISA reaction, and only a small fraction of the total free antibody in the solution remains on the plate (so that the measurement does not significantly affect the equilibrium in the solution).
MST is a biophysical method that detects the affinity of antibody molecules over a wide range of concentrations. It judges the affinity between molecules by measuring the movement of molecules along the infrared laser-induced microscopic temperature gradient. This movement depends on many factors, including the hydrated shell, charge and molecular size. These molecules are initially uniformly distributed in solution and free to diffuse. When the infrared laser is turned on, the unbound molecules usually move out of the heating point. The binding of one molecule to another (such as antibodies and antigens) causes the movement of the entire temperature gradient to change. The movement of the molecule can then be followed by fluorescent labeling to derive affinity parameters for the antibody molecule.
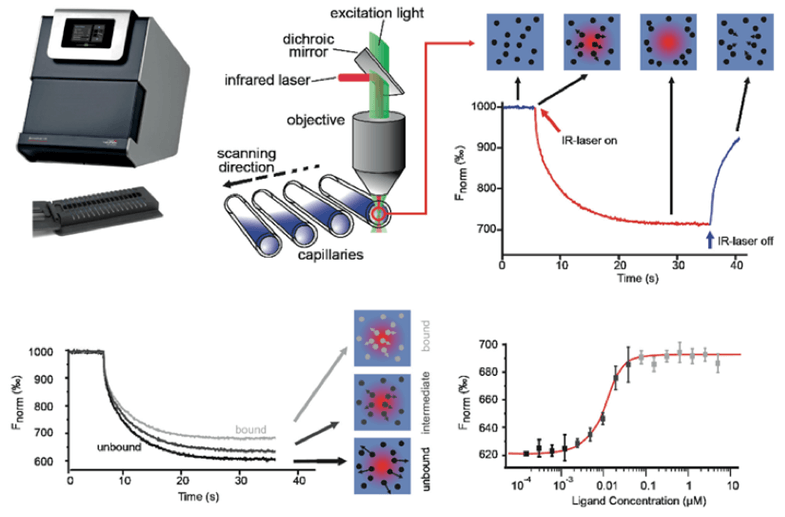 Figure 2. Determination of antibody affinity by Microscale Thermophoresis (MST).
Figure 2. Determination of antibody affinity by Microscale Thermophoresis (MST).
SPR is an optical technique for detecting molecular interactions in which one molecule is immobilized in a metal film and the other molecule is mobile. The combination of these molecules changes the refractive index of the film. Therefore, when polarized light strikes the film, the extinction angle of the light changes and can be monitored by an optical detector (Figure 3).
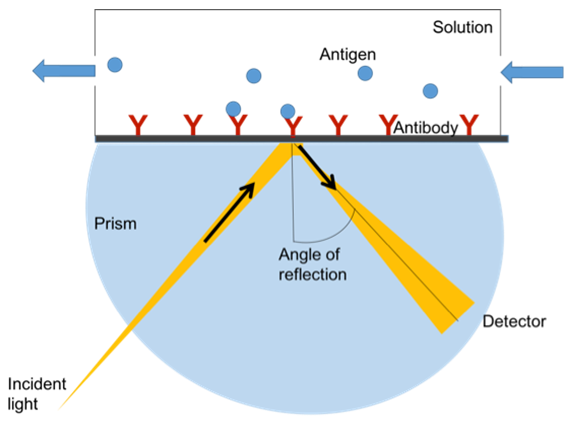 Figure 3. Determination of antibody affinity by Surface Plasmon Resonance (SPR).
Figure 3. Determination of antibody affinity by Surface Plasmon Resonance (SPR).
Finally, verify the reproducibility of the antibody, which is an essential part of antibody detection. A common misconception in antibody testing is to assume that antibodies produce similar results, whether from the same or different batches or from different manufacturers. One of the most shocking examples is David Rim of Yale University. He developed an antibody-based assay to guide the effective treatment of melanoma and is ready to apply the assay to the clinic. However, when he ordered a new set of antibodies from the same company, he could not reproduce the original results and had to give up his work. That is to say, using antibodies from the same supplier over time, even if only the batch is different, will produce different results. Therefore, it is critical for researchers to always test for antibody repeatability. This becomes especially important when using polyclonal antibodies, as products from the same item number from the supplier may mean different antibodies.
In summary, in order to ensure that the antibody meets the specificity and affinity and reproducibility required for its use, it is necessary to perform a secondary verification strictly according to the antibody detection standard. Only in this way can the reliability of the experimental data be guaranteed.