 Figure 1. Western blot experimental steps 1~5. From sample preparation to protein electrophoresis.
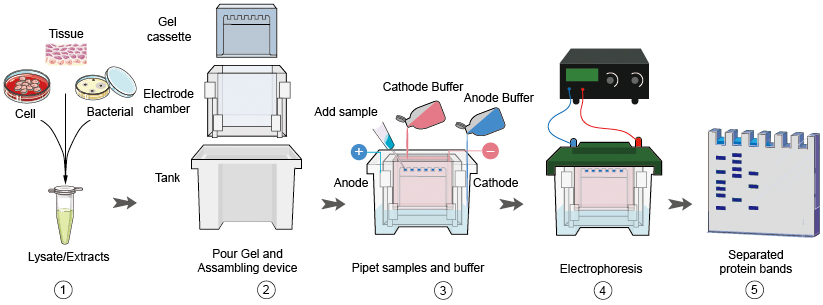
Figure 1. Western blot experimental steps 1~5. From sample preparation to protein electrophoresis.
Sample preparation is the first step and one of the most important steps of western blot. A good sample preparation makes your western blot half success. By understanding the nature of your starting sample and having a clear picture of the information you wish to derive from your Western blotting experiments, the chances of a successful analysis could be insured.
In principle, all sources of protein, from single cells to whole tissues as well as extracellular matrices, biological fluids and proteins secreted in vitro, are open to analysis by Western blotting. Whereas sources such as mammalian cells in suspension are easily disrupted under mild conditions and readily release their proteins, it is more difficult to extract proteins from intact tissues or within solid tumors. Extraction of the proteins from plants, bacteria and fungi are further complicated. Regardless of the source and protein of interest, however, the aim must be to harvest the proteins of interest, while at the same time, obtaining a sufficient yield of proteins at an acceptable level of purity.
General tips of sample preparation:
Be gentle! Stay cool!
Protein extract:
Here we list some recommended sample extract method for several kinds of materials in Table 1.
Table 1. Overview of extraction options for different cells and tissues
| Sample | Typical lysis options |
| Tissue culture | Detergent lysis |
| Cell suspensions | Ultrasonication |
| Most plant and animal tissues | Mechanical homogenization |
| Soft animal tissues and cells | Manual or mechanical homogenization |
| Bacterial and mammalian cells | Freeze/thaw lysis |
| Bacteria, erythrocytes, cultured cells | Osmotic shock lysis |
| Solid tissues and plant cells | Manual grinding with mortar and pestle |
| Cell suspensions, yeast cells | Grinding with abrasive component |
| Bacteria, yeast, plant tissues, fungal cells | Enzymatic digestion |
| Bacteria, yeast, plant cells | Nitrogen cavitation |
| Microorganisms with cell walls | French press |
| Plant tissues, fungal cells | Glass bead milling |
Lysis buffers:
The extracted protein must be dissolved and preserved in lysis buffer to further conduct the following electrophoresis process. There are many recipes for lysis buffers but a few will serve for most western blotting experiments. In brief, they differ in their ability to solubilize proteins, with those containing sodium dodecyl sulfate and other ionic detergents considered to be the harshest and therefore most likely to give the highest yield. However, these detergent can cause protein denature more or less. So if your antibody used cannot recognize denatured proteins, you should avoid the detergent in buffer such as SDS, deoxycholate, Triton X-100 and NP-40 or try to use some relatively mild non-ionic detergents.
Protease and phosphatase inhibitors
Protease inhibitors must be included in lysis buffers to prevent degradation of proteins following the release of endogenous proteases during the process of cell lysis. As soon as lysis occurs, proteolysis, de-phosphorylation and denaturation begin. These events can be slowed down tremendously if samples are kept on ice or at 4°C at all times and appropriate inhibitors are added fresh to the lysis buffer. Cocktails of inhibitors from various suppliers are available, but still you can make your own inhibitor mixture. Here we list some inhibitor formula in Table 2.
Table 2. Inhibitor formula
| Inhibitor | Protease/phosphatase inhibited | Final concentration in lysis buffer | Stock (store at -20°C) |
| Aprotinin | Trypsin, Chymotrypsin, Plasmin | 2 μg/ml | Dilute in water, 10 mg/ml. Do not re-use once defrosted. |
| Leupeptin | Lysosomal | 5-10 μg/ml | Dilute in water. Do not re-use once defrosted. |
| Pepstatin A | Aspartic proteases | 1 μg/ml | Dilute in methanol, 1 mM. |
| PMSF | Serine, Cysteine proteases | 1 mM | Dilute in ethanol. You can re-use the same aliquot. |
| EDTA | Metalloproteases that require Mg2+ and Mn2+ | 5 mM | Dilute in H2O, 0.5 M. Adjust pH to 8.0. |
| EGTA | Metalloproteases that require Ca2+ | 1 mM | Dilute in H2O, 0.5 M. Adjust pH to 8.0. |
| Na Fluoride | Serine/Threonine phosphatases | 5-10 mM | Dilute in water. Do not re-use once defrosted. |
| Na Orthovanadate | Tyrosine phosphatases | 1 mM | Dilute in water. Do not re-use once defrosted. |
Preparation of lysate from cell culture:
Preparation of lysate from tissues:
Protein quantification:
After extracting from cells or tissue lysate, protein samples should be quantification so that to compare the amount of protein from samples run in different lanes within the same gel or between gels and make all the lanes have been loaded with the same total amount of protein. Several spectrophotometric methods are routinely used to determine the concentration of protein in a solution. These include measurement of the intrinsic ultraviolet (UV) absorbance of the protein as well as methods based on a protein-dependent color change, such as the classic, copper-based Lowry assay, the Smith copper/bicinchoninic assay (BCA) and the Bradford dye assay.
Table 3 compare of three different protein quantification methods.
| Introduction | Advantages: | Disadvantages: | |
| The Lowry Method | Relies on the reaction of copper with proteins, but the sample is also incubated with the Folin-Ciocalteu reagent. Reduction of the Folin-Ciocalteu reagent under alkaline conditions results in an intense blue color (heteropolymolybdenum blue) that absorbs at 750 nm. The Lowry method is best used with protein concentrations of 0.01–1.0 mg/mL. | Easy to use Highly reproducible Inexpensive Sensitive and broad linear range | Need to do a standard curve for every assay Timing and mixing of reagents must be precise |
| Bradford Assay | Binding of Coomassie Brillant Blue G-250 to proteins, causes a shift of the dye from red (465 nm) to blue (595 nm) under acidic conditions. It is compatible with more common reagents, although detergents can cause interference. Proteins with a concentration of 20-2000 μg/mL can be measured using the Bradford assay. | Easy to use Sensitive and broad linear range Quick Compatible with many buffers | Need to do a standard curve for every assay Reagent stains cuvettes Often need to dilute samples prior to analysis Depends strongly on amino acid composition |
| Bicinchoninic Acid Assay (BCA) | After reduction of Cu2+ ions, two molecules of BCA chelate with each Cu+ ion resulting in formation of an intense purple color that absorbs at 560 nm. BCA is as sensitive as the Lowry method and works well with protein concentrations from 0.5 μg/mL to 1.5 mg/mL. | Easy to use Highly reproducible Inexpensive Sensitive and broad linear range | Need to do a standard curve for every assay Color continues to develop over time, but is stable for measurement after 30 minutes at 37°C |
Once the concentration of each sample have been determined, you can freeze them at -20°C or -80°C for later use or prepare for loading onto a gel or other usage.
Polyacrylamide gels are inert, crosslinked structures. The pore sizes in these gels are similar to the molecular radius of many proteins. As molecules are forced through the gel in an electric field, larger molecules are retarded by the gel more than smaller molecules. When separated on a polyacrylamide gel, the procedure is abbreviated as SDS-PAGE (for Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis). The technique is a standard means for separating proteins according to their molecular weight. Polyacrylamide gels are formed from the polymerization of two compounds; acrylamide and N, N-methylenebis-acrylamide (Bis, for short). Bis is a cross-linking agent for the gels. The polymerization is initiated by the addition of ammonium persulfate along with either DMAP or TEMED. The gels are neutral, hydrophilic, three-dimensional networks of long hydrocarbons crosslinked by methylene groups. The separation of molecules within a gel is determined by the relative size of the pores formed within the gel. The pore size of a gel is determined by two factors: the total amount of acrylamide present and the amount of cross-linker. As the percentage of acrylamide increases, the pore size decreases. With cross-linking 5% gives the smallest pore size. Any increase or decrease in cross-linking percentage increases or decreases the pore size. Gels are designated as percent solutions and will have two necessary parameters. The total acrylamide is given as a percentage (w/v) of the acrylamide plus the bis-acrylamide. The average pore size is determined by the percentage of the amount of cross linker and total amount of acrylamide used. Polyacrylamide is used to separate most proteins, ranging in molecular weight from Mr >5000 to<200,000.
Table 4 Recommended acrylamide concentration for protein target within defined size ranges.
| Target protein size range (Mr) | Recommended acrylamide concentration |
| 36 000 to 205 000 | 5% |
| 24 000 to 205 000 | 7.5% |
| 14 000 to 205 000 | 10% |
| 14 000 to 66 000 | 12.5% |
| 14 000 to 45 000 | 15% |
The percentage of polyacrylamide used in the gel along with the buffer system will influence the mobility of the proteins through the gel as current is applied. The expected size of the target protein can be used to select the best gel/buffer system to achieve optimal separation and resolution (Figure 2).
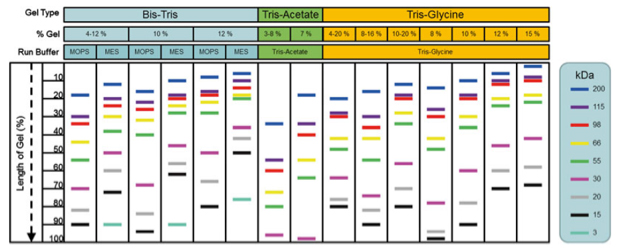 Figure 2. Migration of protein by molecular weight varies by gel type, concentration, and running buffer. Use the estimated migration patterns provided here to assist in the selection of gel type and running buffer based on the predicted MW of the target protein and the desired separation
Figure 2. Migration of protein by molecular weight varies by gel type, concentration, and running buffer. Use the estimated migration patterns provided here to assist in the selection of gel type and running buffer based on the predicted MW of the target protein and the desired separation
The SDS-PAGE system we used is usually a discontinuous system. It consists of two different gel: the stacking gel and the separating/resolving gel (Figure 3.). Stacking gel usually with low pH (6.8) and Acr Bis concentration (4%), that makes the stacking gel have higher porosity and could hardly influence the movement of proteins. Separating gel with higher pH (8.8) and Acr Bis concentration (12.5%). Use a gel comb to form sample loading cells on stacking gel. Load the sample in the cells and switch on the power. By initially running the samples through a lower density stacking gel, proteins are concentrated in a matter of minutes into a thin starting zone by the time the sample contents reach the resolving gel by a process known as isotachophoresis. The interface between the two gel densities may thus be regarded as the starting line for all the proteins in each well and on entering the resolving gel, the proteins begin to separate according to size. A reference recipe of these two kind of gel preparation is shown in Table 5 and Table 6.
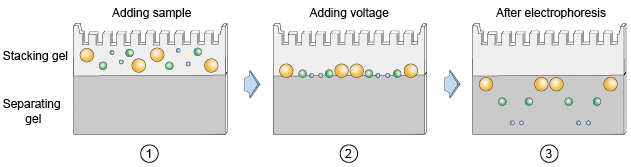 Figure 3. The schematic diagram of stacking gel & separating gel.
Figure 3. The schematic diagram of stacking gel & separating gel.
Table 5. Reference recipe of 5% stacking gel in different volume.
| 5% Stacking gel | Volume of different component needed in different volume of gel (mL) | |||||||
| 1 | 2 | 3 | 4 | 5 | 6 | 8 | 10 | |
| ddH2O(mL) | 0.68 | 1.4 | 2.1 | 2.7 | 3.4 | 4.1 | 5.5 | 6.8 |
| 30% Acr Bis (mL) | 0.17 | 0.33 | 0.5 | 0.67 | 0.83 | 1 | 1.3 | 1.7 |
| 1M Tris (mL) pH=6.8 | 0.13 | 0.25 | 0.38 | 0.5 | 0.63 | 0.75 | 1 | 1.25 |
| 10% SDS (μL) | 10 | 20 | 30 | 40 | 50 | 60 | 80 | 100 |
| 10% APS* (μL) | 10 | 20 | 30 | 40 | 50 | 60 | 80 | 100 |
| TEMED* (μL) | 1 | 2 | 3 | 4 | 5 | 6 | 8 | 10 |
| *, APS: Ammonium Persulfate ; TEMED: Tetramethylethylenediamine; SDS: Sodium dodecyl sulfate | ||||||||
Table 6. Reference recipe of 5mL separating gel in different percentage.
| 5mL Separating gel | Volume of different component needed in different percentage of gel (%) | ||||||||||||
| 4% | 5% | 6% | 7% | 8% | 9% | 10% | 11% | 12% | 13% | 14% | 15% | 16% | |
| ddH2O(mL) | 3 | 2.8 | 2.65 | 2.5 | 2.3 | 2.15 | 2 | 1.8 | 1.65 | 1.5 | 1.3 | 1.15 | 1 |
| 30% Acr Bis (mL) | 0.65 | 0.85 | 1 | 1.15 | 1.35 | 1.5 | 1.65 | 1.85 | 2 | 2.15 | 2.35 | 2.5 | 2.65 |
| 1.5 M Tris (mL) pH=8.8 | 1.25 | ||||||||||||
| 10% SDS (μL) | 50 | ||||||||||||
| 10% APS* (μL) | 50μL | ||||||||||||
| TEMED* (μL) | 3μL | ||||||||||||
| *, APS: Ammonium Persulfate; TEMED: Tetramethylethylenediamine; SDS: Sodium dodecyl sulfate | |||||||||||||
Learn more about western blot, please see Next Section.